Press Release5 Minute ReadDec | 4 | 2019
Mass General researchers uncover early adherence step “hidden in plain sight” in intestinal transit of Shigella pathogen

Key Takeaways
- The bacterial pathogen Shigella, often spread through contaminated food or water, is a leading cause of mortality in both children and older adults in the developing world
- No effective vaccine has been developed, and the pathogen has acquired resistance to many antibiotics
- The recent discovery of an early adherence step in the infection cycle could provide a new therapeutic target or even a new method for vaccine development.
Christina Faherty, PhDWe discovered what had been hidden in plain sight before—the gene expression profiles that enabled Shigella to initiate this early step in infection by attaching to the epithelial tissue of the host.”
Mucosal Immunology and Biology Research Center
Massachusetts General Hospital
Boston, MA – The bacterial pathogen Shigella, often spread through contaminated food or water, is a leading cause of mortality in both children and older adults in the developing world. Although scientists have been studying Shigella for decades, no effective vaccine has been developed, and the pathogen has acquired resistance to many antibiotics. The recent discovery of an early adherence step in the infection cycle by researchers at Massachusetts General Hospital (MGH) could provide a new therapeutic target or even a new method for vaccine development.
As it moves through the digestive system, Shigella traverses the small intestine and subsequently infects the large intestine, causing cramping, diarrhea and dehydration in the disease called shigellosis. “We wanted to determine how Shigella makes its first contact with epithelial cells in the early stages of disease development,” says Christina Faherty, PhD, investigator in the Mucosal Immunology and Biology Research Center at MGH and senior author on the study published in mSphere. “Because of certain gene sequence annotations, and the way that Shigella appeared following growth in standard laboratory media, it was believed that Shigella strains do not produce fimbriae or other adherence factors.” Fimbriae are short hair-like fibers that bacterial cells use to adhere to individual epithelial cells to instigate infection.
The work of Faherty and the research team has uncovered evidence of fimbriae that aid adherence to epithelial cells, an important step in the start of a shigellosis infection. “We mimicked the conditions that Shigella would face in its journey through the small intestine by adding bile salts and glucose to laboratory media,” says Faherty. “With this method, we discovered what had been hidden in plain sight before—the gene expression profiles that enabled Shigella to initiate this early step in infection by attaching to the epithelial tissue of the host.”
Researchers at the Mucosal Immunology and Biology Research Center at MGH performed comprehensive microscopy and genetic analyses of Shigella to determine its subsequent steps after leaving the stomach. Their results demonstrate that “at least three structural genes facilitate S. flexneri (strain) 2457T adherence for epithelial cell contact and biofilm formation.” In other words, their findings contradict the current hypothesis that critical components in the gene clusters are unable to produce fimbriae or other adherence factors.
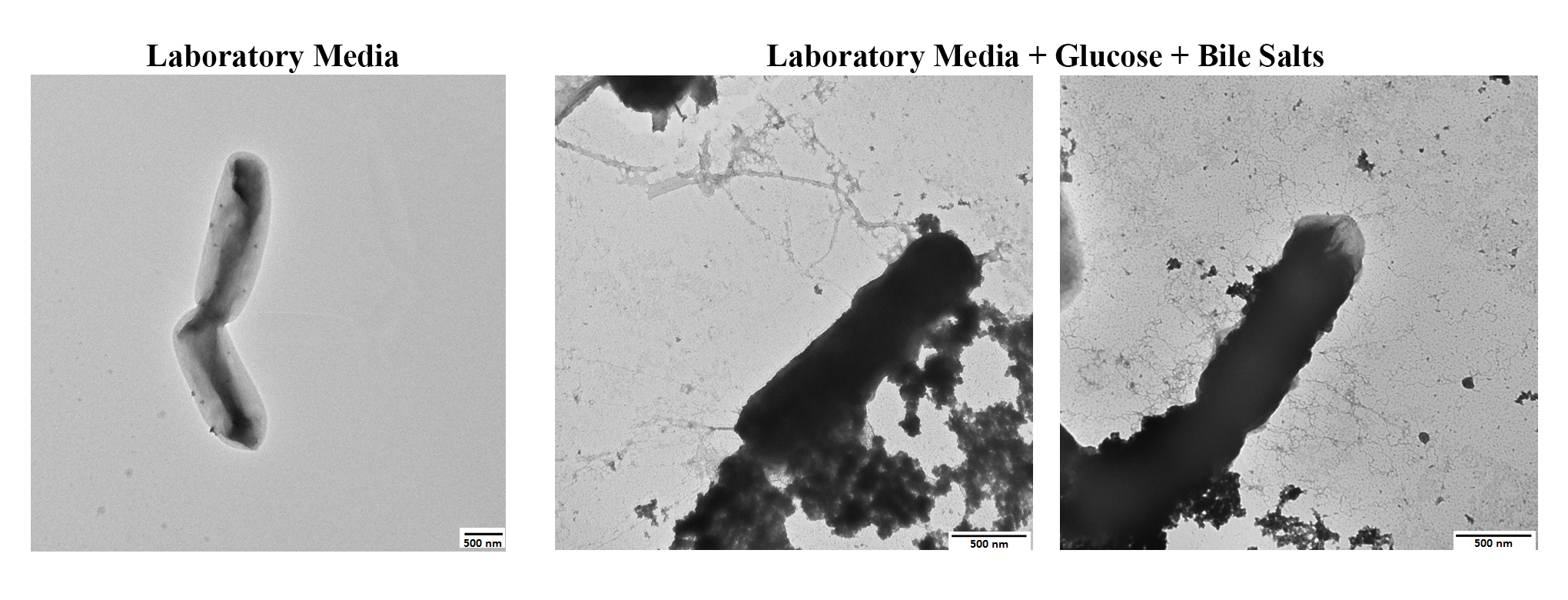

Transmission electron microscopy of Shigella flexneri strain 2457T grown in laboratory media (left) or grown in laboratory media supplemented with glucose and bile salts (center and right). Three types of adherence factors expressed by the bacteria can be visualized. (Image panels reproduced with permission from mSphere.)
In earlier research, Faherty and colleagues determined that exposure to bile salts resulted in the formation of biofilms, a protective coating of bacterial communities. Faherty hypothesizes that this coating enables the pathogen to survive the harsh conditions of the small intestine to successfully enter the colon. Since biofilm formation requires adherence factors, and since bacterial cells dispersed from the biofilm adhere better to epithelial cells, the next step by the group was to investigate adherence factor expression under these conditions. This next step was indeed controversial given the hypotheses that Shigella does not produce adherence structures; yet, the comprehensive analyses provided strong evidence to the contrary.
Co-author Rachael Chanin notes that the group’s most recent study confirms their earlier analyses that the “in vivo-like” conditions facilitated biofilm formation and adherence to epithelial cells through fimbriae attachment. “One of the main challenges in studying Shigella is the lack of animal models that faithfully recapitulate human disease,” says Chanin. “Although there have been elegant and thorough studies of what happens when the pathogen enters colonic epithelial cells, we did not understand what happens during transit through the digestive system or how the bacterium approaches or interacts with host cells prior to entry. Our work begins to address these questions and underlines the importance of in vivo-like culture methods. It also shows that these methods may influence our experimental results—whether intentionally or unintentionally.”

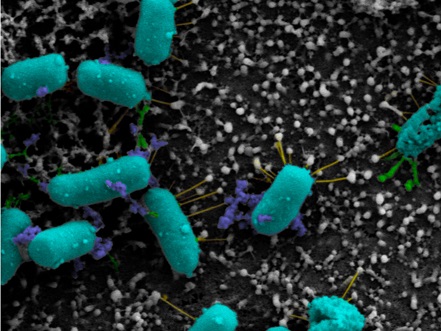
Scanning electron microscopy of Shigella flexneri strain 2457T adhering to the apical surface of a colonic epithelial cell from the organoid model. Pseudo-coloring was performed to visualize the bacterial components: teal (bacterial cells), green (thicker adherence factors), yellow (thinner adherence factors) and blue (electron-dense aggregates). (Image reproduced with permission from mSphere.)
After the promising results from their bile salts and glucose laboratory model, the researchers added another component to their adherence analysis—a human intestinal organoid. The “mini-gut,” created from stem cells isolated from intestinal tissue, represents a model of the human intestinal epithelium. Working with a mini-gut of the ascending colon, the researchers discovered the Shigella adherence structures making initial contact with epithelial cells. “We think these adherence factors used in the intestinal organoid model replicate the contact made with the epithelial cells in the colon in the initial stages of shigellosis,” says Faherty.The researchers plan to deepen future investigations through in-depth analyses of adherence factor production and secretion and the study of components found in different Shigella strains. “As a field, we are starting to understand how the signals in the gut change during inflammation, and in the future, we can leverage this knowledge to better understand pathogenesis and better model disease,” says Chanin.
About the Massachusetts General Hospital
Massachusetts General Hospital, founded in 1811, is the original and largest teaching hospital of Harvard Medical School. The MGH Research Institute conducts the largest hospital-based research program in the nation, with an annual research budget of more than $1 billion and comprises more than 8,500 researchers working across more than 30 institutes, centers and departments. In August 2019 the MGH was once again named #2 in the nation by U.S. News & World Report in its list of "America’s Best Hospitals."
