Research Summary
The Ellisen laboratory has a broad interest in how genetic abnormalities in breast cancer and related malignancies influence tumor biology and how that biology can, in turn, be exploited to therapeutic advantage. We address these questions through basic research studies of key cancer hallmarks. including DNA repair defects through BRCA1/2 and related pathways and transcriptional reprogramming through the p53 gene family. Supporting and complementing these studies are sophisticated analyses of patient-derived precancerous and cancerous tissues. Recent innovative tissue-based studies have led to our discovery of novel cancer drivers and have provided a unique window into early cancer pathogenesis, intratumoral heterogeneity, and therapeutic resistance. Our discoveries in the basic laboratory and through human tumor analysis are being applied in ongoing clinical trials that seek to identify predictive markers of response to innovative therapeutics for breast and other cancers. Our ability to work at the interface of basic tumor biology and therapeutic application is strongly supported by our network of collaborators and by the research and clinical infrastructure of the Mass General Brigham Cancer Institute.
Research Projects
Novel drivers of aggressive breast cancer subtypes
Our work employing advanced tumor molecular diagnostics has revealed gene fusions as novel drivers of an aggressive breast cancer subset. In triple-negative breast cancer (TNBC), extensive intratumoral heterogeneity is itself a driver that we have characterized through single-cell genomic and transcriptomic analysis, leading to our discovery of unanticipated drug resistance mechanisms with immediate therapeutic implications. Of particular interest is resistance to novel Antibody Drug Conjugates (ADCs) that are transforming cancer therapy. Unraveling the complex nature of ADC resistance is a long-term goal of ours that touches every aspect of tumor biology and will have major clinical impact. Our longstanding work on the biology of TNBC is supported by the institution-wide Triple Negative Breast Cancer Program, which integrates basic research, translational and clinical studies together with human tumor propagation and high-throughput drug screening, all focused on overcoming drug resistance and improving outcomes for patients with TNBC.
BRCA1/2, hereditary cancer predisposition and prevention
Germline mutations in the DNA repair genes BRCA1 and BRCA2 confer dramatically elevated risk of cancers of the breast, ovary, and pancreas, yet the precise pathogenesis of BRCA1/2-associated cancer remains to be elucidated. Together with an international team of collaborators we are carrying out systematic studies of early events that give rise to these cancers, in part through detailed molecular analysis of normal and precancerous tissues from BRCA1/2 mutation carriers. Defining the altered signaling and early cooperating events in this context is likely to reveal new markers of breast cancer predisposition and new targets for prevention. For example, our published single-cell genome analysis has revealed extensive chromosomal damage in BRCAmutant breast tissues that precedes any histological abnormalities. This seminal finding implies the existence of early cellular defects and associated vulnerabilities that could be exploited for cancer prevention in this setting.
The p53 family network in cancer biology and therapy
As a transcription factor and key nodal point for integrating cellular stress responses, the p53 tumor suppressor controls diverse cellular processes, including cell cycle progression, survival, and metabolism. Through analysis of two p53-related genes, p63 and p73, we and others have defined a functional network including a tissue specific role for p63 as the enforcer of an epigenetically controlled stem/progenitor state. Tumor-selective deregulation of p63 and associated chromatin remodeling factors reprogram the transcriptome to inhibit differentiation and promote immune evasion. These findings likely explain why p63 is over-expressed in a large variety of epithelial tumors, particularly squamous cell and breast carcinomas. Collectively, this work serves as a paradigm for the analysis of transcriptional reprogramming in cancer.
Research Positions
Postdoctoral Research Fellow (1)
One Postdoctoral Research Fellow position is currently available. The candidate must have recently received a PhD degree in the biological sciences, and be highly motivated and well versed in basic molecular biology and biochemical techniques. The Fellow will have simultaneous academic appointments at the Massachusetts General Hospital and Harvard Medical School. The position provides a rich intellectual environment with full integration into the large research communities of the Mass General and Harvard. The laboratory studies fundamental mechanisms of tumorigenesis and their associated therapeutic implications in breast and other cancers. Topics of major interest include the p53 family of transcription factors, hereditary breast cancer and molecular genetics. We have identified fundamental mechanisms of the p53 family members in tumorigenesis (Cancer Cell 2017; 31:35) and normal development (Dev Cell 2014; 30:151), and have discovered novel genetic breast cancer drivers (Cancer Discovery 2017). We recently uncovered a new stress-induced tumor suppressor pathway involving regulation of mTOR activity (Nat Comm 2015; 6:7014), and revealed for the first time frequent gene regulatory mutations in breast cancer (Nature 2017; 547:55). Our ability to work at the interface of basic tumor biology, genetics and therapeutic application is strongly supported by the research and clinical infrastructure of the Mass General Brigham Cancer Institute.
To apply, please email a brief cover letter and CV to:
Leif W. Ellisen, MD, PhD
Mass General Brigham Cancer Institute
CPZN 4204
185 Cambridge Street
Boston, MA 02114
Email: lellisen@mgh.harvard.edu
Publications
View a list of publications by researchers at the Ellisen Laboratory
Selected Publications
Coates JT, Sun S, Leshchiner I, Thimmiah N, Martin EE, McLoughlin D, Danysh BP, Slowik K, Jacobs RA, Rhrissorrakrai K, Utro F, Levovitz C, Denault E, Walmsley CS, Kambadakone A, Stone JR, Isakoff SJ, Parida L, Juric D, Getz G, Bardia A, and Ellisen LW. Parallel genomic alterations of antigen and payload targets mediate polyclonal acquired clinical resistance to sacituzumab govitecan in triple-negative breast cancer. Cancer Discovery. 2021 11:1-10.
Koh SB, Ellisen LW. Immune activation and evolution through chemotherapy plus checkpoint blockade in triplenegative breast cancer. Cancer Cell. 2021 Dec 13;39(12):1562-1564.
Qiao S, Koh SB, Vivekanandan V, Salunke D, Patra KC, Zaganjor E, Ross K, Mizukami Y, Jeanfavre S, Chen A, Mino-Kenudson M, Ramaswamy S, Clish C, Haigis M, Bardeesy N, and Ellisen LW. REDD1 loss reprograms lipid metabolism to drive progression of RAS-mutant tumors. Genes Dev. 2020 Jun 1;34(11-12):751-766.
Karaayvaz M, Silberman RE, Langenbucher A, Saladi SV, Ross KN, Zarcaro E, Desmond A, Yildirim M, Vivekanandan V, Ravichandran H, Mylavagnanam R, Specht MC, Ramaswamy S, Lawrence M, Amon A, Ellisen LW. Aneuploidy and a deregulated DNA damage response suggest haploinsufficiency in breast tissues of BRCA2 mutation carriers. Science Advances 2020;6:5.
Karaayvaz M, Cristea S, Gillespie SM, Patel AP, Mylvaganam R, Luo CC, Specht MC, Bernstein BE, Michor F, and Ellisen LW. Unravelling subclonal heterogeneity and aggressive disease states in TNBC through single-cell RNA-seq. Nature Communications. 2018 (In Press).
Matissek KJ, Onozato ML, Sun S, Zheng Z, Schultz A, Lee J, Patel K, Jerevall PL, Saladi SV, MacLeay A, Tavallai M, Badovinac-Crnjevic T, Barrios C, Be?e N, Chan A, Chavarri-Guerra Y, Debiasi M, Demirdogen E, Egeli U, Gökgöz S, Gomez H, Liedke P, Tasdelen I, Tolunay S, Werutsky G, St Louis J, Horick N, Finkelstein DM, Le LP, Bardia A, Goss PE, Sgroi DC, Iafrate AJ, Ellisen LW. Expressed Gene Fusions as Frequent Drivers of Poor Outcomes in Hormone Receptor Positive Breast Cancer. Cancer Discovery2018 8(3):336-353.
Saladi, SV, Ross K, Karaayvaz M, Tata PR, Mou H, Rajagopal J, Ramaswamy S, and Ellisen LW. ACTL6A is co-Amplified with p63 in Squamous Cell Carcinoma to Drive YAP Activation, Regenerative Proliferation and Poor Prognosis. Cancer Cell 2017 31:35-49.
Research Image
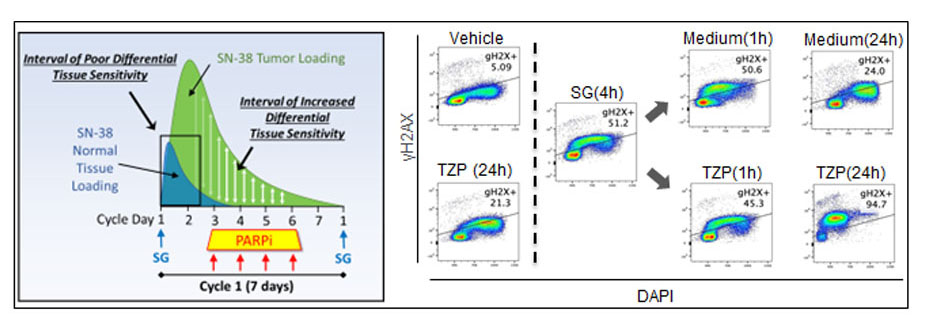
The schematic at left demonstrates that tumor-selective delivery of cytotoxic SN-38 via the Antibody-Drug Conjugate Sacituzumab govitecan (SG) allows normal cells to rapidly clear the drug, while sequential PARP inhibitor (PARPi) treatment is toxic to tumor cells with residual SN-38. At right, flow cytometry plots showing SG induces DNA damage (g-H2AX) that is rapidly repaired following washout (Medium) but progresses to lethal damage in the presence of PARP inhibitor Talazoparib (TZP). The concept of sequential SG/TZP dosing was successfully applied in our clinical trial for advanced breast cancer (Bardia, Ellisen et al, Clin. Cancer. Res. 2024).




