Inaugural Krantz Awards Recipient
2023 Quantum Award: Targeting transcription factors to open doors to new cancer therapies
Team: Liron Bar-Peled, PhD, Michael Lawrence, PhD and Chris Ott, PhD.
Learn more about the team's project and the Krantz Awards
Research Summary
Research in the Bar-Peled laboratory sits at the interface of cellular metabolism and signal transduction and focuses on understanding how cancer cells respond to altered metabolic states. Rapidly proliferating cancer cells are characterized by increased production of toxic metabolic byproducts known as reactive oxygen species (ROS) that at high levels potently block cancer cell growth. To neutralize high ROS levels, cancer cells activate the NRF2 pathway, which governs the cellular antioxidant response. While the NRF2 pathway is critical for cancer growth, the molecular mechanisms by which this pathway functions and provides cancer cells with a proliferative advantage remain poorly understood. By combining frontier molecular, chemical and proteomic approaches, research in our lab has revealed that NRF2 establishes a unique cellular environment that protects critical proteins required for cancer cell growth from inactivation by ROS. Our studies indicate that these ROS-regulated proteins are highly targetable by small molecule inhibitors and may be exploited to develop chemical tools to inactivate these dependencies in cancers.
Research Projects
Cancer cells display remarkable plasticity allowing them to adapt to ever changing environments. A key feature of this plasticity is their ability to rewire core metabolic networks to provide a steady source of energy and building blocks needed for rapid growth. This demand for energy produces byproducts, including ROS that alters the function of proteins, DNA and lipids, and if left unchecked, results in oxidative stress and impairs cancer cell viability. To counter a rise in oxidative stress, cells activate the NRF2 transcription factor leading to the expression of a vast network of antioxidant and detoxification genes that restore redox homeostasis. Multiple cancer cells, including ~30% of non-small cell lung cancers (NSCLCs) activate NRF2 through the genetic disruption of its negative regulator KEAP1. Despite its clear importance in cancer cell proliferation, we know remarkably little about how the NRF2/KEAP1 pathway functions within cancer cells or how ROS modification of proteins alters their function. Our longterm goal is to understand how cancer cells sense and respond to ROS and to pharmacologically modulate these pathways in cancers where they are deregulated.
Redox control pathways in Lung Cancer
Our recent studies focus on how the intracellular environment generated by NRF2 in NSCLCs is required for cancer cell proliferation. By employing a chemical proteomics platform (isoTOP-ABPP) that identifies changes in cysteine reactivity mediated by ROS, we demonstrated that NRF2 is required for the protection of dozens of proteins from ROS modification. We found that silencing NRF2 in NSCLCs reduced the reactivity of the catalytic cysteine of the glycolytic enzyme GAPDH without changing GAPDH protein abundance. Concomitant knockdown of NRF2 significantly reduced GAPDH enzyme activity and glycolytic flux, a metabolic pathway required to fuel cancer cell proliferation. These results illustrate how NRF2 can regulate enzyme and pathway activity, not through direct transcriptional control, but rather by fostering a favorable redox environment required for proper enzyme function. Current studies in our lab seek to elucidate how other proteins are post-translationally regulated by NRF2 and feedback into this pathway. To address these questions, we are studying the function of ROS-regulated sites on proteins as well as the identifying reactive metabolites that modify them.
Druggable co-dependencies
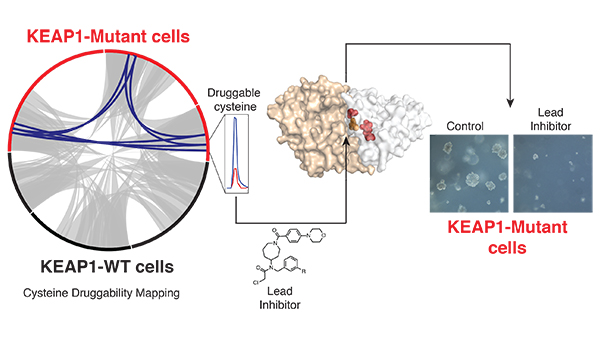
Our investigations suggest that the cellular state created by NRF2 may be exploited to develop inhibitors targeting proteins whose expression and function are stimulated by this environment. Because of their importance to protein function, cysteines are targeted by multiple clinically approved inhibitors. To identify pharmacological targets of the NRF2 pathway, we use powerful chemical proteomic platforms (cysteine druggability mapping) to identify the landscape of protein druggability (e.g. ligand-protein interactions) in genetically defined lung cancers. Our studies reveal that multiple proteins, including the orphan nuclear receptor NR0B1, are exclusively druggable in KEAP1-mutant, NRF2- activated cells. By developing a small molecule inhibitor that disrupts NR0B1 protein interactions we show that NR0B1 functions as a critical signaling node within the NRF2 pathway to support its proproliferative transcriptional output required for anchorage-independent growth. Recently we uncovered that cysteine residues that are sensitive to ROS modification are highly targetable by covalent inhibitors. Our current studies suggest that these sites may be exploited to develop inhibitors that target proteins required for the proliferation of NRF2- activated cancers.
Ongoing projects:
- Determine how cancer proteomes respond to changes in the intracellular redox environment
- Elucidate the role of NRF2-regulated reactive metabolites on protein function
- Decipher how cells adapt to anchorage-independent growth
- Identify druggable transcriptional dependencies in genetically-defined cancers
Research Positions
The Bar-Peled (BP) Lab is now recruiting new lab members. Our research is interdisciplinary and as such we look for researchers who are ambitious, motivated, and unafraid to go in the directions that their research takes them. Success in research comes not only from persistence, creativity, and focus but also mentorship. We are committed to providing an exciting research environment that will allow you to reach your maximum scientific potential and a mentoring environment that will help you pursue your broader goals. Please send a cover letter, your CV and a list of three references to Dr. Liron Bar-Peled at lbar-peled@mgh.harvard.edu.
The following positions are open:
Postdoctoral fellows: Postdoctoral fellows interested in combining molecular, chemical and proteomic approaches to address fundamental questions in metabolic signaling are encouraged to apply. A background in cellular or chemical biology is highly desirable. Applicants with a pure synthetic chemistry background who wish to use their chemical expertise to interrogate biological problems are very welcome as well. Email Liron with a Cover letter your CV and contact information for three references.
Research Technicians/Associates: We are seeking highly motivated candidates who enjoy conducting research and involving themselves in the dynamic life of the laboratory. She or he will have the opportunity to become involved with all the steps of research being completed by our group and work closely with Dr. Bar-Peled. The experience gained in the lab will be perfect for those seeking entrance to graduate or medical school. College degree with a preference in biology, molecular biology, or biochemistry. Candidates with a computational background are also welcome to apply. 1-3 years of independent research experience is required. The ideal candidate will be detail-oriented, organized, and able to work independently as well as part of a team in a challenging environment. Excellent communication and organizational skills are necessary. Email Liron with a Cover letter your CV and contact information for three references. Your cover letter should indicate your career goals and employment timeline, your laboratory skill set and previous research experience.
Principal Responsibilities for Research Associates:
- Prepares basic solutions and performs base-level procedures as assigned (i.e. – pipetting, cell and tissue culture, etc.)
- Prepares proteomic samples
- Conducts mammalian tissue culture experiments
- Conducts molecular biology experiments (cloning, western blotting, etc..)
- Maintains laboratory notebook
- Understands and applies basic scientific techniques
- Conducts analysis of results
- Sets up and prepares routine experiments as directed
- Prepares lab reagents, chemicals, instruments and equipment
- Preforms independent literature searches
- Assists with organizing materials for publication or presentation
- Maintains and orders supplies
Postdoctoral fellows interested in combining molecular, chemical and proteomic approaches to address fundamental questions in metabolic signaling are encouraged to apply. A background in cellular or chemical biology is highly desirable. Applicants with a pure synthetic chemistry background who wish to use their chemical expertise to interrogate biological problems are very welcome as well. Email Liron with a Cover letter your CV and contact information for three references.
Doctoral students and masters students interested in helping us understand how cells respond to altered metabolic states using cutting edge molecular and chemical platforms should apply. Doctoral students interested in a rotation should email Liron to set a up a time to meet. Current masters students hoping to work in the lab as a visiting student (6-12 month duration) to complete your thesis are also welcome. We are looking for enthusiastic and curious individuals who share our passions.
We are seeking highly motivated candidates who enjoy conducting research and involving themselves in the dynamic life of the laboratory. She or he will have the opportunity to become involved with all the steps of research being completed by our group. The experience gained in the lab will be useful for pursuing a career as a researcher or physician. College degree with a preference in biology, molecular biology, or biochemistry. Candidates with a computational background are also welcome to apply. 1-3 years of independent research experience is required. The ideal candidate will be detail-oriented, organized, and able to work independently as well as part of a team in a challenging environment. Excellent communication and organizational skills are necessary. Email Liron with a Cover letter your CV and contact information for three references. Your cover letter should indicate your career goals and employment timeline, your laboratory skill set and previous research experience.
Undergraduate students, who have completed relevant coursework and wish to begin conducting biomedical research as part of their thesis project are welcome to apply. Indicate how much time you plan to spend in the lab and what your goals are.
The Bar-Peled lab is part of the Krantz Family Center for Cancer Research at Massachusetts General Hospital and is located in the Charlestown Navy Yard. We are also part of the Department of Medicine at Harvard Medical School. We can be contacted here.
Publications
Selected Publications
Takahashi M†, Chong HB, Zhang S, Yang TY, Lazarov MJ, Harry S, Maynard M, Hilbert B, White RD, Murrey HE, Tsou CC, Vordermark K, Assaad J, Gohar M, Dürr BR,… Oh E, Fisher DE, Maheswaran S, Haber DA, Boland GM, Sade-Feldman M, Jenkins RW, Hata AN, Bardeesy NM, Suvà ML, Martin BR, Liau BB, Ott CJ, Rivera MN, Lawrence MS†, Bar-Peled L†. DrugMap: A quantitative pan-cancer analysis of cysteine ligandability. Cell. 2024 Apr 17:S0092-8674(24)00318-0.
Weiss-Sadan T, Ge M†, Hayashi M, Gohar M, Yao CH, de Groot A, Harry S, Carlin A, Fischer H, Shi L, Wei TY, Adelmann CH, Wolf K, Vornbäumen T, Dürr BR, Takahashi M, Richter M, Zhang J, Yang TY, Vijay V, Fisher DE, Hata AN Haigis MC, Mostoslavsky R, Bardeesy N, Papagiannakopoulos T, Bar-Peled L†. NRF2 activation induces NADHreductive stress, providing a metabolic vulnerability in lung cancer. Cell Metab. 2023 Apr 4;35(4):722.
Zhang J†, Simpson CM, Berner J, Chong HB, Fang J, Ordulu Z, Weiss- Sadan T, Possemato AP, Harry S, Takahashi M, Yang TY, Richter M, Patel H, Smith AE, Carlin AD, Hubertus de Groot AF, Wolf K, Shi L, Wei TY, Dürr BR, Chen NJ, Vornbäumen T, Wichmann NO, Mahamdeh MS, Pooladanda V, Matoba Y, Kumar S, Kim E, Bouberhan S, Oliva E, Rueda BR, Soberman RJ, Bardeesy N, Liau BB, Lawrence M, Stokes MP, Beausoleil SA, Bar-Peled L†. Systematic identification of anticancer drug targets reveals a nucleus-tomitochondria ROS- sensing pathway. Cell. 2023 May 25;186(11):2361-2379
Bar-Peled L*†, Kemper EK*, Suciu RM, Vinogradova EV, Backus KM, Horning BD, Paul TA, Ichu TA, Svensson RU, Olucha J, Chang MW, Kok BP, Zhu Z, Ihle N, Dix MM, Hayward M, Jiang P, Saez E, Shaw RJ, and Cravatt BF†. (2019) Chemical Proteomics Identifies Druggable Vulnerabilities in a Genetically Defined Cancer. Cell, 171(3), 696–709.e23.
Bar-Peled L*, Chantranupong L*, Cherniack AD, Chen WW, Ottina KA, Grabiner BC, Spear ED, Carter SL, Meyerson ML, and Sabatini DM. (2013). A tumor suppressor complex with GAP activity for the Rag GTPases that signal amino acid sufficiency to mTORC1. Science 340: 1100-1106.
*These authors contributed equally to this work
†Co-corresponding authors